Overview of protein synthesis
The pathway of protein synthesis is called translation because the language of the nucleotide course on the mRNA is translated into the language of the amino acid sequence.
The genetic information which is stored in the chromosome are finally translated into proteins and are the end products of most information pathway.
The translation process requires a genetic code through which the information contained in the nucleic acid sequence that expressed to produce a specific sequence of amino acids in the body.
The Genetic Code:
The genetic code is a glossary that defines the correspondence between a sequence of nucleotides bases in the body & the sequence of amino acids in humans.
Each individual word in the code is composed of 3 specific nucleotide bases, these genetic words are called codons.
Codons
Codons are presented in the mRNA that language of adenine (A), guanine (G), cytosine (C), uracil (U). Their nucleotide sequence is every time in a 5’ – 3’ direction.
There are 4 nucleotide bases that are implemented to produce 3 base codons. Therefore there are 64 different types of combinations of bases taking 3 at a time.
There are four important codons which are the following below :
- Start codon: AUG (methionine)
- Stop codon: UAG, UGA, UAA.
Stop codons do not code for amino acids in the body, so they are terminate the codon.
Characteristics of genetic code:
- Specificity: The code is specific (unambiguous), i.e. a particular codon most of the time codes for the same amino acid.
- Universality: They are universal in nature, their specificity has been maintained from an early stage of evolution with only a little difference except in the case of the mitochondria UGA codes for tryptophan.
- Degeneracy: Occasionally called redundant although each codon that corresponds to a single amino acid, an amino acid given that may have more than one codon, for example, leucine, serine, arginine, etc have 6 different codons, however methionine and tryptophan have one codon.
- Nonoverlapping and commaless genetic code.
Consequences of altering the nucleotide sequence:
- Silent mutation:
- The codon that containing the altered base may code for the same amino acid.
- For example, if the serine codon UCA is given a different third base—U—to turn out to be UCU, it still codes for serine.
- Missense mutation:
- The codon that containing the altered base may code for a different amino acid.
- For example, if the serine codon UCA is given a different first base—C—to turn out to be CCA, it will code for proline.
- Nonsense mutation:
- The codon that containing the altered base may become a termination codon.
- For example, if the serine codon UCA is given a different second (2nd) base—A—to turn out to be UAA, the new codon causes a cessation of translation at that point.
- Other mutations:
- These can alter or change the amount or structure of the protein produced by the translation process in the body.
- Trinucleotide repeat expansion:
- A sequence of 3 bases that are repeated in tandem will turn out to be amplified in number so that too several copies of the triplet occur in the body.
- For example, amplification of the CAG codon leads to the insertion of several extra glutamine residues in the huntingtin protein, which causes Huntington’s disease.
- Splice site mutations:
- Mutations at splice locations can alter or change the way in which introns are removed from pre-mRNA molecules, which producing aberrant proteins.
- Frame-shift mutations:
- If one or two nucleotides are either removed from or added to the coding area of a message sequence, a frame-shift mutation occurs, & the reading frame is changed.
- This can result in a product with a radically dissimilar or different amino acid sequence, or a truncated product due to the formation of a termination codon.
- Trinucleotide repeat expansion:
- These can alter or change the amount or structure of the protein produced by the translation process in the body.
Components required for Translation (protein synthesis):
- Aminoacid.
- Transfer RNA.
- Aminoacyl tRNA synthetase.
- Messenger RNA.
- Ribosome.
- Protein factors.
- ATP and GTP an energy sources.
Wobble hypothesis:
There are 64 probable codons. For translation, each of these codons requisite a tRNA molecule with a complementary anticodon.
If each tRNA molecule paired with its complementary mRNA codon that using Watson-Crick base pairing, then 64 different types of tRNA molecule would be required.
Since most organisms have fewer than 45 different species of tRNA, some tRNA species must pair with several than one codons.
The anticodons in some tRNAs involve the nucleotide inosinate (designated I), which contains the abnormal base hypoxanthine. Inosinate can form hydrogen bonds with three different types of nucleotides (U, C, & A).

So in 1966, Francis Crick initiated (proposed) the Wobble hypothesis to account for this.
- The first 2 bases of an mRNA codon consistently form strong Watson-Crick base pairs with the corresponding bases of the tRNA anticodon & give out to most of the coding specificity.
- The first base of the anticodon dictates the number of codons identify by the tRNA.
- When the 1st base of the anticodon is C or A, base pairing is specific & only one codon is identified by that tRNA.
- When the 1st base is U or G, that binding is less specific 7 two different types of codons may be read.
- When inosine (I) is the 1st (wobble) nucleotide of an anticodon, 3 different types of codons(U, C &A) can be identified as the largest number for any tRNA.
- When an amino acid is specified by several dissimilar codons, the codons that vary in either of the first two bases require different tRNAs.
- A minimum of 32 tRNAs is needed to translate all 61 different codons (31 to encode the amino acids & one for initiation).
- Since the 3rd base of the codon sets (pairs) loosely with the anticodon, it allows rapid dissociation of tRNA from its codon throughout protein synthesis.
- If all 3 bases of a codon involved in strong Watson-Crick pairing with the anticodon, tRNAs would separate too slowly and this would gravely limit the rate of protein synthesis.
- Codon-anticodon interactions equilibrium the requirements for accuracy & speed.
Steps in Protein Synthesis
The procedure (process) of protein synthesis translates the three (3) letter alphabet of nucleotide sequence on mRNA into a 20 letter alphabet of amino acid in the body.
An important differentiation between prokaryotes & eukaryotes. In prokaryotes, translation & transcription are coupled i.e. translation beginning before transcription is accomplished, this is because of the lack of nuclear membrane in prokaryotes. There are actual steps in protein synthesis are the following;
- Initiation
- Elongation
- Termination
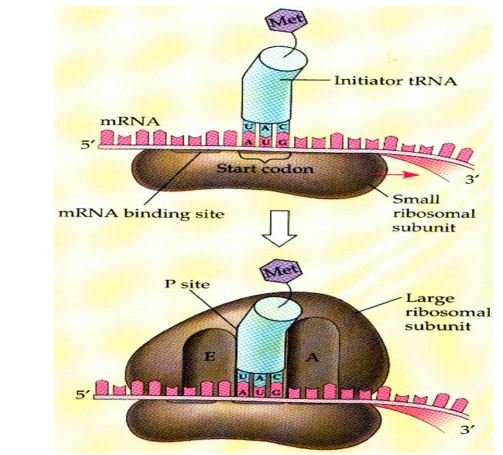
Initiation
- 1st gathering or assembly of a component of translation that occurs which includes – 2 ribosomal subunits, & mRNA to be translated, aminoacyl tRNA, GTP, initiation factors in the body.
- Initiation factors in prokaryotes such as IF 1, IF 2, IF 3.
- Initiation factors in eukaryotes such as more than 10 – eIF.
- There are two mechanisms by which the ribosome recognizes AUG that initiate the translation process.
- First mechanism:
- Prokaryotes:
- Shine Dalgarno sequence situated 6-10 bases upstream of initiating AUG codon base pair with 16S RNA consequently facilitating the positioning of small (30S) ribosomal subunit in mRNA in near proximity to AUG.

- Shine Dalgarno sequence situated 6-10 bases upstream of initiating AUG codon base pair with 16S RNA consequently facilitating the positioning of small (30S) ribosomal subunit in mRNA in near proximity to AUG.
- Prokaryotes:
- The second mechanism (Initiation codon):
- Initiating AUG that recognized by a special initiator tRNA in this mechanism.
- Recognization facilitated by IF2GTP in prokaryotes microorganism and eIF-2GTP + other eIF in eukaryotes organisms.
- The charged initiator tRNA directly enters the P site.
- In bacteria & mitochondria, initiator tRNA carries N–formylated methionine.
- In eukaryotes, initiator tRNA carries methionine that is not formylated.
- The huge ribosomal subunit then joins the complex & a functional ribosome which is formed with charged initiating tRNA in P-site.
Elongation
It involves or participates in the addition of amino acids to the carboxyl end of the growing chain. This ribosome moves from 5’ – 3’ extremity.
Then aminoacyl tRNA whose codon appears next on the mRNA template carrying the 2nd amino acid moves to site A.
This is facilitated by, in prokaryotes includes EF-Tu GTP and EF-Ts, in eukaryotes organisms, includes EF-1α GTP and EF-1βγ.
Formyl methionine which is carried by tRNA in the P site is then connected to the amino acid which is carried by tRNA that has just entered the A site, by a peptide bond which is catalyzed by peptidyl transferase (ribozyme) enzyme.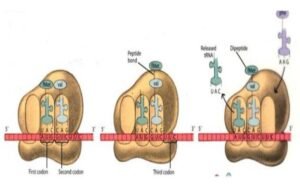
Translocation
The ribosome then advances three nucleotides towards the 3’ extremity of mRNA known as translocation.
For translocation: in prokaryotes that requires EF-G-GTP and eukaryotes that requires EF-2-GTP. Translocation causes following in the body,
- Movement of uncharged tRNA from P site to E site.
- Movement of peptidyl tRNA from the A site to the P site.
This process occurs repeatedly until the termination codon is encountered.
Termination
It occurs when one of the 3 termination codons moves into the A site. These codons are recognized or notified by the release factor (RF).
Which causes hydrolysis of the bond between peptide & tRNA at the P site that releasing the nascent protein from the ribosome in the body.
In prokaryotes;
- RF1 recognizes or identifies UAA and UAG.
- RF2 recognizes or identifies UGA and UAA.
- RF-3-GTP releases or identifies RF1 and RF2.
- In eukaryotes organisms, only one release factor eRF recognizes or identifies all three termination codons.
Once the nascent (newly formed) protein is released other components such as ribosomal subunits, mRNA, tRNA, & protein factors are also released and recycled or reused for the synthesis of another polypeptide chain in the body.
[embeddoc url=”https://notesmed.com/wp-content/uploads/2020/08/protein-synthesis.pdf” download=”all” cache=”off”]